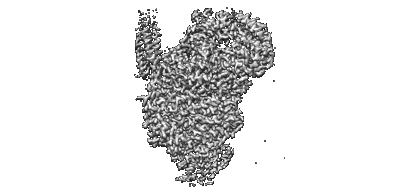
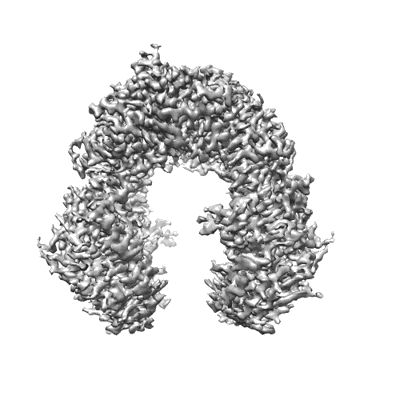Cryo-EM structure of the human parainfluenza virus hPIV3 L-P polymerase in dimeric form
Resolution: 2.7 Å
EM Method: Single-particle
Fitted PDBs: 8kdb
Q-score: 0.577
Xie J, Ouizougun-Oubari M, Wang L, Zhai G, Wu D, Lin Z, Wang M, Ludeke B, Yan X, Nilsson T, Gao L, Huang X, Fearns R, Chen S
Nat Commun (2024) 15 pp. 3163-3163 [ DOI: doi:10.1038/s41467-024-47470-7 Pubmed: 38605025 ]
- Zinc ion (65 Da, Ligand)
- Rna-directed rna polymerase l (260 kDa, Protein from Human respirovirus 3)
- Magnesium ion (24 Da, Ligand)
- Human parainfluenza virus hpiv3 l-p polymerase in dimeric form (Complex from Human respirovirus 3)
- Phosphoprotein (68 kDa, Protein from Human respirovirus 3)
Cryo-EM structure of the human parainfluenza virus hPIV3 L-P polymerase in monomeric form
Resolution: 3.3 Å
EM Method: Single-particle
Fitted PDBs: 8kdc
Q-score: 0.506
Xie J, Ouizougun-Oubari M, Wang L, Zhai G, Wu D, Lin Z, Wang M, Ludeke B, Yan X, Nilsson T, Gao L, Huang X, Fearns R, Chen S
Nat Commun (2024) 15 pp. 3163-3163 [ DOI: doi:10.1038/s41467-024-47470-7 Pubmed: 38605025 ]
- Zinc ion (65 Da, Ligand)
- Rna-directed rna polymerase l (260 kDa, Protein from Human respirovirus 3)
- Magnesium ion (24 Da, Ligand)
- Phosphoprotein (68 kDa, Protein from Human respirovirus 3)
- Human parainfluenza virus hpiv3 l-p polymerase in monomeric form (Complex from Human respirovirus 3)
Structure of Gabija AB complex
Resolution: 2.79 Å
EM Method: Single-particle
Fitted PDBs: 8tjy
Q-score: 0.534
Yang XY, Shen Z, Xie J, Greenwald J, Marathe I, Lin Q, Xie WJ, Wysocki VH, Fu TM
Nat Struct Mol Biol (2024) [ DOI: doi:10.1038/s41594-024-01283-w Pubmed: 38627580 ]
- Endonuclease gaja (67 kDa, Protein from Bacillus cereus)
- Gabija protein gajb (57 kDa, Protein from Bacillus cereus)
- Tetramer of gabija protein a (Complex from Bacillus cereus)
Structure of Gabija AB complex
Resolution: 3.23 Å
EM Method: Single-particle
Fitted PDBs: 8tk0
Q-score: 0.496
Yang XY, Shen Z, Xie J, Greenwald J, Marathe I, Lin Q, Xie WJ, Wysocki VH, Fu TM
Nat Struct Mol Biol (2024) [ DOI: doi:10.1038/s41594-024-01283-w Pubmed: 38627580 ]
- Endonuclease gaja (67 kDa, Protein from Bacillus cereus)
- Tetramer of gabija protein a (Complex from Bacillus cereus)
Structure of Gabija AB complex 1
Resolution: 2.98 Å
EM Method: Single-particle
Fitted PDBs: 8tk1
Q-score: 0.453
Yang XY, Shen Z, Xie J, Greenwald J, Marathe I, Lin Q, Xie WJ, Wysocki VH, Fu TM
Nat Struct Mol Biol (2024) [ DOI: doi:10.1038/s41594-024-01283-w Pubmed: 38627580 ]
- Tetramer of gabija protein ab (4:4) (Complex from Bacillus cereus)
- Endonuclease gaja (67 kDa, Protein from Bacillus cereus)
- Gabija protein gajb (57 kDa, Protein from Bacillus cereus)
Structure of E. coli PtuA hexamer
Resolution: 2.93 Å
EM Method: Single-particle
Fitted PDBs: 8sux
Q-score: 0.5
Li Y, Shen Z, Zhang M, Yang XY, Cleary SP, Xie J, Marathe IA, Kostelic M, Greenwald J, Rish AD, Wysocki VH, Chen C, Chen Q, Fu TM, Yu Y
Nat Struct Mol Biol (2024) 31 pp. 413-423 [ DOI: doi:10.1038/s41594-023-01172-8 Pubmed: 38177683 ]
- Ptua (53 kDa, Protein from Escherichia coli)
- Ptua (Complex from Escherichia coli)
- Adenosine-5'-triphosphate (507 Da, Ligand)
Resolution: 2.6 Å
EM Method: Single-particle
Fitted PDBs: 8ee4
Q-score: 0.576
Li Y, Shen Z, Zhang M, Yang XY, Cleary SP, Xie J, Marathe IA, Kostelic M, Greenwald J, Rish AD, Wysocki VH, Chen C, Chen Q, Fu TM, Yu Y
Nat Struct Mol Biol (2024) 31 pp. 413-423 [ DOI: doi:10.1038/s41594-023-01172-8 Pubmed: 38177683 ]
- Ptua (53 kDa, Protein from Escherichia coli)
- 6 ptua form as a complex (Complex from Escherichia coli)
- Adenosine-5'-triphosphate (507 Da, Ligand)
Structure of focused PtuA(dimer) and PtuB(monomer) complex
Resolution: 2.72 Å
EM Method: Single-particle
Fitted PDBs: 8ee7
Q-score: 0.609
Li Y, Shen Z, Zhang M, Yang XY, Cleary SP, Xie J, Marathe IA, Kostelic M, Greenwald J, Rish AD, Wysocki VH, Chen C, Chen Q, Fu TM, Yu Y
Nat Struct Mol Biol (2024) 31 pp. 413-423 [ DOI: doi:10.1038/s41594-023-01172-8 Pubmed: 38177683 ]
- Ptua (53 kDa, Protein from Escherichia coli)
- 6 ptua form as a complex (Complex from Escherichia coli)
- Ptub (28 kDa, Protein from Escherichia coli)
- Adenosine-5'-triphosphate (507 Da, Ligand)
Structure of E.coli Septu (PtuAB) complex
Resolution: 2.6 Å
EM Method: Single-particle
Fitted PDBs: 8eea
Q-score: 0.603
Li Y, Shen Z, Zhang M, Yang XY, Cleary SP, Xie J, Marathe IA, Kostelic M, Greenwald J, Rish AD, Wysocki VH, Chen C, Chen Q, Fu TM, Yu Y
Nat Struct Mol Biol (2024) 31 pp. 413-423 [ DOI: doi:10.1038/s41594-023-01172-8 Pubmed: 38177683 ]
- 6 ptua and 2 ptub form as a complex (Complex from Escherichia coli)
- Ptua (53 kDa, Protein from Escherichia coli)
- Ptub (28 kDa, Protein from Escherichia coli)
- Adenosine-5'-triphosphate (507 Da, Ligand)
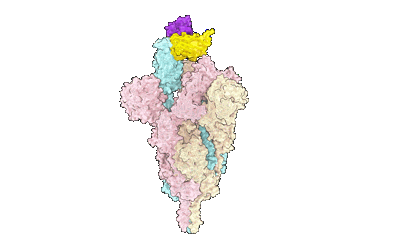
SARS-CoV-2 Omicron variant spike in complex with Fab 9A8 (State 1)
Resolution: 3.4 Å
EM Method: Single-particle
Fitted PDBs: 7wt7
Q-score: 0.385
Liu S, Jia Z, Nie J, Liang Z, Xie J, Wang L, Zhang L, Wang X, Wang Y, Huang W
Cell Discov (2022) 8 pp. 81-81 [ Pubmed: 35977939 DOI: doi:10.1038/s41421-022-00443-w ]
- Spike glycoprotein (141 kDa, Protein from Severe acute respiratory syndrome coronavirus 2)
- Heavy chain of fab 9a8 (12 kDa, Protein from Homo sapiens)
- Light chain of fab 9a8 (11 kDa, Protein from Homo sapiens)
- Sars-cov-2 omicron variant spike in complex with fab 9a8 (state 1) (Complex from Severe acute respiratory syndrome coronavirus 2)
- 2-acetamido-2-deoxy-beta-d-glucopyranose (221 Da, Ligand)